FLAT - Flowgram Alignment Tool
Supplementary Information
This web page contains supplementary information for the paper "A
probabilistic method for small RNA flowgram matching,"
Vladimir Vacic,
Hailing Jin,
Jian-Kang Zhu
and Stefano Lonardi,
Pacific Symposium on Biocomputing, PSB'08, 13:75-86. (2008)

 slides
slides
Supplementary Figure 1. Distributions of signal strengths for
three 454 pyrosequencing datasets: A) A. thaliana (50 million
flows, small RNA discovery); B) H. sapiens (38.4 million flows,
small RNA discovery); C) C. bifermentans (188.9 million flows,
whole-genome sequencing). The overlaps between Gaussians for
different polynucleotide lengths are responsible for over-calling or
under-calling the lengths of incorporated nucleotide runs.
It is of interest to observe that in the H. sapiens dataset, the
means of Gaussians for length 5 poly-C,G,T have been skewed towards
lower values. Also, peaks for 7 poly-A,T are higher than the peaks for
6 poly-A,T, which implies that in the sample there are more 7-mers
than 6-mers - which is clearly not possible.
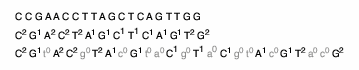
Supplementary Figure 2. Flowspace encoding of the sequence
CCGAACCTTAGCTCAGTTGG: the second line shows run-length encoding
(RLE) of the sequence, and the third line shows insertions of dummy
negative flows (gray lower case letters). The flowspace encoding is
the output of an ideal sequencer, which does not make mistakes in
terms of lengths of polynucleotides.
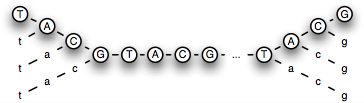
Supplementary Figure 3. Combinations of negative flows which may
flank a subsequence (the capital letters signify appropriately padded
run length encoding of the sequence database, and lower case letters
signify negative flows). Here all 16 combinations are allowed:
TACG...TACG, tACG...TACG, taCG...TACG, tacG...TACG, ..., taCG...TACg,
taCG...TAcg, taCG...Tacg, ..., tacg...tacg).
Acknowledgments
V.V. and S.L. were supported in part by NSF CAREER IIS-0447773, and NSF
DBI-0321756. H.J. was supported in part by NSF CAREER MCB-0642843 and
AES-CE Research Allocation Award PPA-7517H.
The authors would like to thank Shou-Wei Ding (Department of Plant Pathology,
UC Riverside) and Sarjeet Gill (Cell Biology and Entomology) for kindly
providing the additional pyrosequencing data, and Thomas Girke (Botany
and Plant Biology) and Christian Shelton (Computer Science and Engineering)
for useful discussions.
|



